WarpHelix — AI Biomedical Research Agent Platform
Advancing life science research at warp speed
Biomedical Research Faces an Efficiency Crisis
Over 3 million biomedical papers are published globally each year, and public databases hold petabytes of omics data. Yet most researchers cannot effectively leverage these resources — not because they lack scientific intuition, but because they lack the tools and skills.
🔴 Three Core Pain Points
1. Severe Bioinformatics Talent Shortage
- The global bioinformatics talent gap continues to widen
- A qualified bioinformatics engineer commands $50K-120K/year; small labs simply can't afford one
- Even at large institutions, bioinformatics teams serve dozens of research groups — queueing for analysis is the norm
- Result: "Wet lab" experiments are done, but data analysis becomes the bottleneck
2. Fragmented Tools, Steep Learning Curves
- Genomic analysis requires BLAST, BWA, GATK, samtools...
- Single-cell analysis requires Scanpy, Seurat, CellRanger...
- Drug discovery requires RDKit, AutoDock, ADMET tools...
- Each tool has different installation, data formats, and parameters
- Result: Environment setup alone takes days; the real science gets shelved
3. Non-Reproducible Analysis Pipelines
- Manual operations are error-prone, parameters rely on "experience"
- Different analysts may reach different conclusions from the same data
- Result: Wasted time, wasted funding, slower research progress
The biggest bottleneck in biomedical research isn't lack of data — it's lack of people who can analyze it.
How WarpHelix Solves These Problems
WarpHelix's core philosophy: Give every researcher a 24/7 "super postdoc".
No programming required. No environment setup. No learning dozens of tools. Describe your research question in natural language, and WarpHelix handles the entire pipeline from planning to execution to reporting.
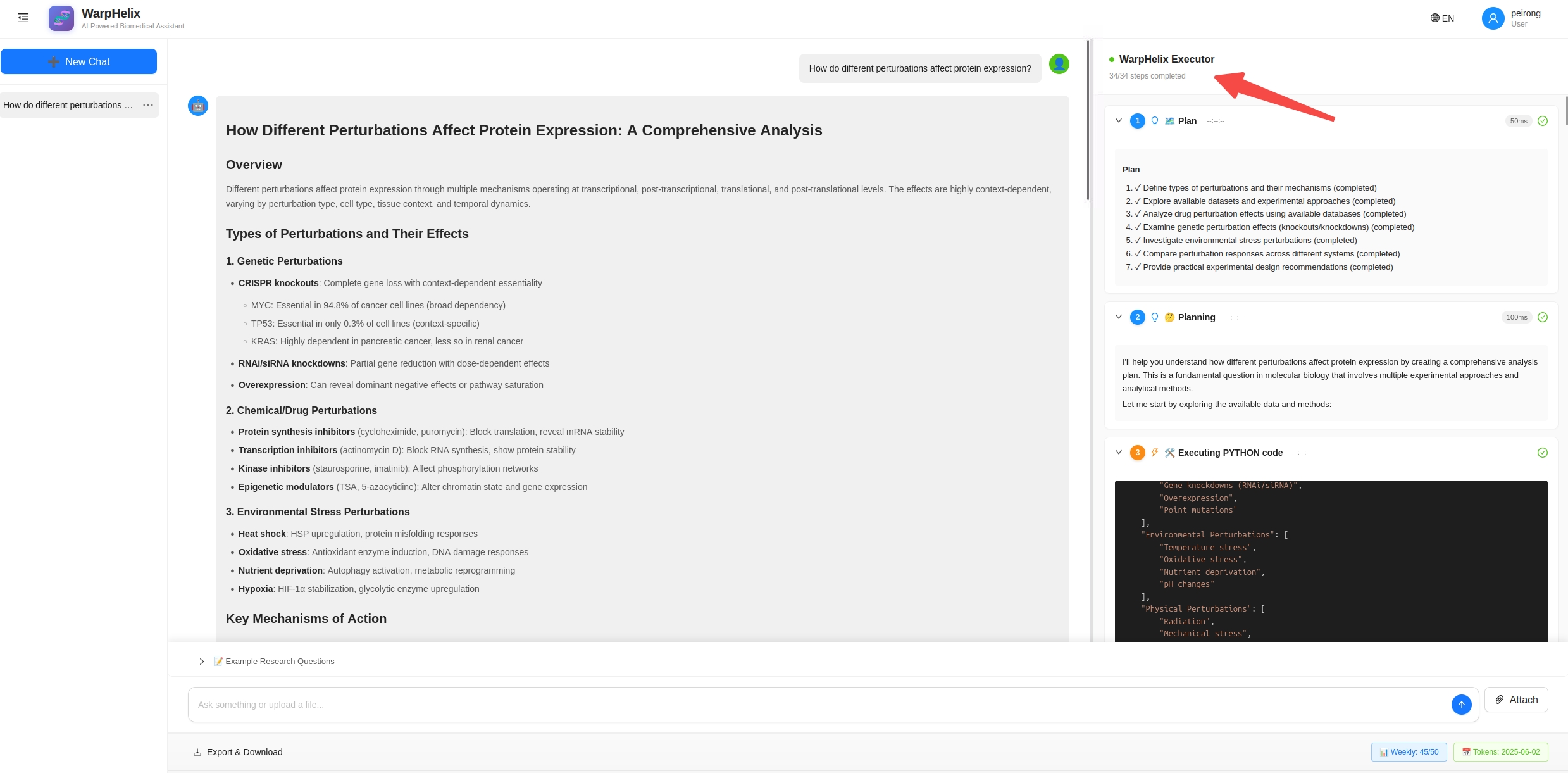
🧠 Core Architecture: Think → Execute → Observe → Solution
Built on Stanford University's Biomni framework, WarpHelix deeply integrates LLM reasoning + retrieval-augmented planning + automated code execution:
Key difference: WarpHelix isn't a "Q&A chatbot" — it's an AI agent that actually does the analysis. It doesn't just tell you what to do; it does it for you.
Why Can WarpHelix Do This?
🔬 150+ Professional Tools, Ready to Use
WarpHelix systematically mines and integrates 150+ professional tools from thousands of biomedical publications — this is Biomni's core innovation: the Action Discovery mechanism.
| Analysis Area | Representative Tools | Typical Tasks |
|---|---|---|
| Sequence Analysis | BLAST, BWA, samtools | Alignment, variant calling |
| Genomics | BEDTools, liftOver, VCF tools | Interval operations, coordinate conversion |
| Single-Cell | Scanpy, AnnData | scRNA-seq analysis, cell annotation |
| Protein | AlphaFold, STRING | Structure prediction, interaction networks |
| Drug Discovery | RDKit, AutoDock, TDC | Molecular docking, ADMET prediction |
| Literature | PubMed, Semantic Scholar | Cross-database search, trend analysis |
🌐 25 Biomedical Subfields Covered
From genomics to cancer genomics, from molecular cloning to rare disease diagnosis — WarpHelix is the most broadly capable biomedical AI agent available.
Comparison: With vs. Without WarpHelix
| Scenario | Traditional Approach | With WarpHelix |
|---|---|---|
| Genetic variant analysis | Manual ClinVar/gnomAD queries, write scripts → 2-3 days | One question, auto-analysis → 5 minutes |
| Single-cell RNA-seq | Setup env, install packages, tune parameters → 1-2 weeks | Upload data, describe needs → 15 minutes |
| Drug ADMET prediction | Install RDKit/TDC, write scripts → 2-3 days | Provide SMILES, get predictions → 3 minutes |
| Protein docking | Configure AutoDock, prepare files → 3-5 days | Specify target protein → 10 minutes |
Enterprise Features
Beyond the open-source Biomni framework, WarpHelix adds production-grade capabilities:
- 👥 Multi-User Management — Complete authentication, roles, and permissions
- 💰 Usage Metering — Precise token tracking with quota management
- 🔧 Multi-Model Support — AWS Bedrock (Claude) + DashScope (Qwen), hot-swappable
- 📁 Smart File Management — Auto-classification, agent environment mounting
- 📤 Full Audit Trail — One-click export of complete analysis sessions
🔒 Data Security: Private Deployment
Biomedical research data is highly sensitive. WarpHelix ensures security at the architectural level:
- Strict user data isolation — Each user's data is independent and inaccessible cross-user
- Research data never used for training — Your data is never used to train AI models
- Full private deployment — All data stays on your servers
- One-click deployment — AWS / Alibaba Cloud / on-premises
👉 View Technical Architecture →
Use Cases
Case 1: Genetic Variant Interpretation — Data to Report in 5 Minutes
Researcher asks: "Analyze these genetic variants, focusing on Parkinson's disease associations."
WarpHelix automatically: Parses VCF → queries ClinVar/gnomAD → retrieves literature → assesses pathogenicity (ACMG) → generates report with visualizations.
Case 2: Single-Cell RNA-seq Full Pipeline
Researcher asks: "Analyze this Perturb-seq dataset, identify key perturbation effects."
WarpHelix automatically: Loads AnnData → QC/normalization → PCA/UMAP → cell annotation → differential expression → pathway enrichment → full report.
Case 3: Drug Developability Assessment
Researcher asks: "Evaluate this compound: CC(=O)Oc1ccccc1C(=O)O"
WarpHelix automatically: Parses SMILES → molecular descriptors → ADMET prediction → Lipinski evaluation → optimization suggestions.
Technology Foundation
WarpHelix is built on Stanford University's Biomni — the first general-purpose biomedical AI agent framework. WarpHelix transforms it from a research prototype into a production-ready enterprise platform.
Contact Us
- 📧 Email: peirongw@foxmail.com
- 🌐 Website: website.autoinfra.cn
- 🏢 Company: Warp Drive Internet Technology Co., Ltd.
WarpHelix — Making AI a super-assistant for every life science researcher
